*-omics
2016-08-12 — 2020-01-22
Wherein the Mapping of Proteomes, Genomes and Phenomes Is Presented as Networks, and Statistical Inference of Control Pathways Using Model Selection and False‑discovery Considerations Is Examined.
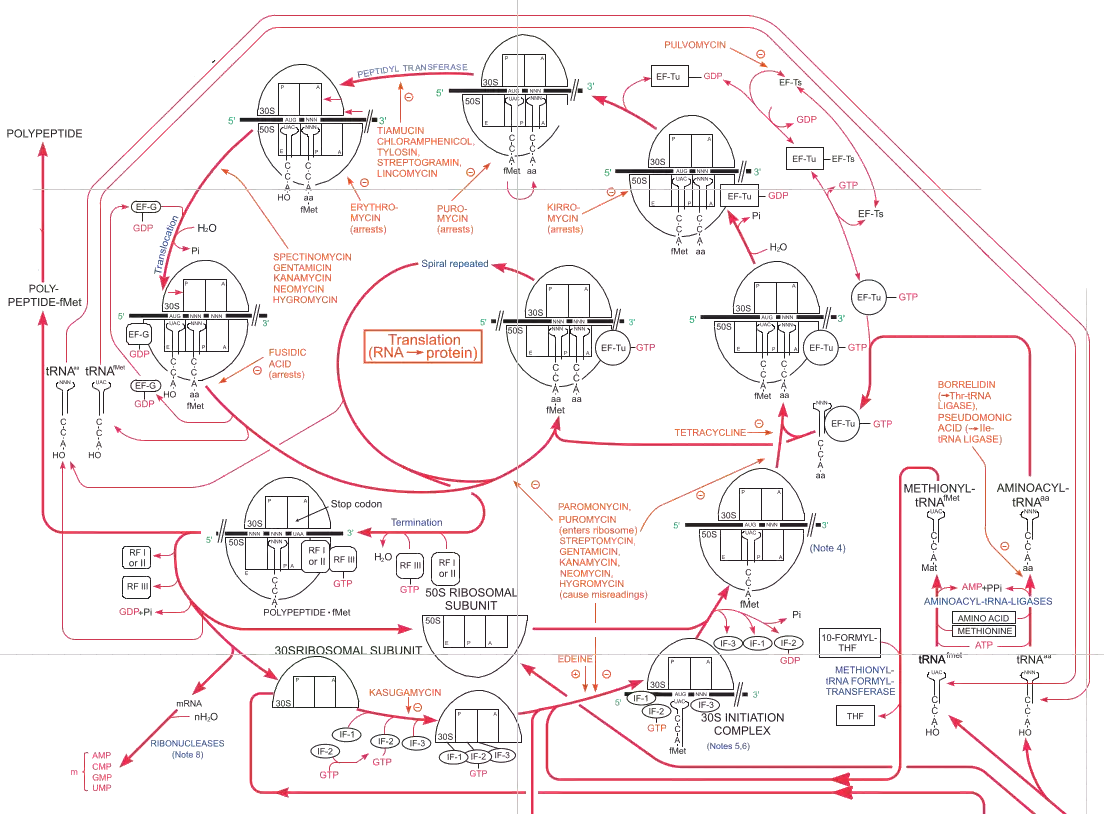
I do not truly understand the Roche biochemical pathways poster.
Proteomics, genomics, phenomics, connectomics. Understanding and inferring networks of control in living systems using statistics generates lots of interesting problems at the nexus of various other statistical problems, like model selection, false discovery rates, causal graphs, and so on.
Of course, there is a deep learning angle.
Is nilearn any good?
1 Incoming
-
By analysing medical text and extracting biomedical entities and relations from the entire history of published medical science, Xyla can facilitate better real-world evidence-based clinical decision support and help make clinical research—such as research into new treatments, including de novo drug design as well as the repurposing of existing drugs—smarter and faster. In so doing, Xyla is fulfilling its mission of organising the world’s medical knowledge and making it more useful.
-
Absci is a generative AI drug creation company. Our mission is simple: create better biologics for patients, faster.