Random-forest-like methods
An optimally-weighted average of randomly stopped clocks is never far from wrong.
2015-09-23 — 2021-06-16
Wherein Ensemble Procedures Composed of Many Weak Decision-Tree Learners Are Treated, Their Aptitude for Tabular Data With Minimal Preprocessing and Apparent Self-Regularizing Behavior Is Recorded, and Implementations Such as XGBoost and LightGBM Are Surveyed
Doubling down on ensemble methods; mixing predictions from many weak learners (typically decision trees) to get strong learners. Boosting, bagging, and other weak-learner ensembles.
There are many flavours of random-forest-like learning systems. The rule of thumb seems to be “Fast to train, fast to use. Gets you results. May not get you answers.” In that regard, they resemble neural networks, but from the previous hype cycle.
Reasons for popularity:
- Decision trees can easily be applied to pretty arbitrary data, so input preprocessing can be minimal; no need to tokenize text or normalize features.
- These methods are, in a certain sense, self-regularizing, so you can skip (certain) hyperparameter tuning.
- There is some kind of tractable asymptotic performance analysis available for some, apparently?
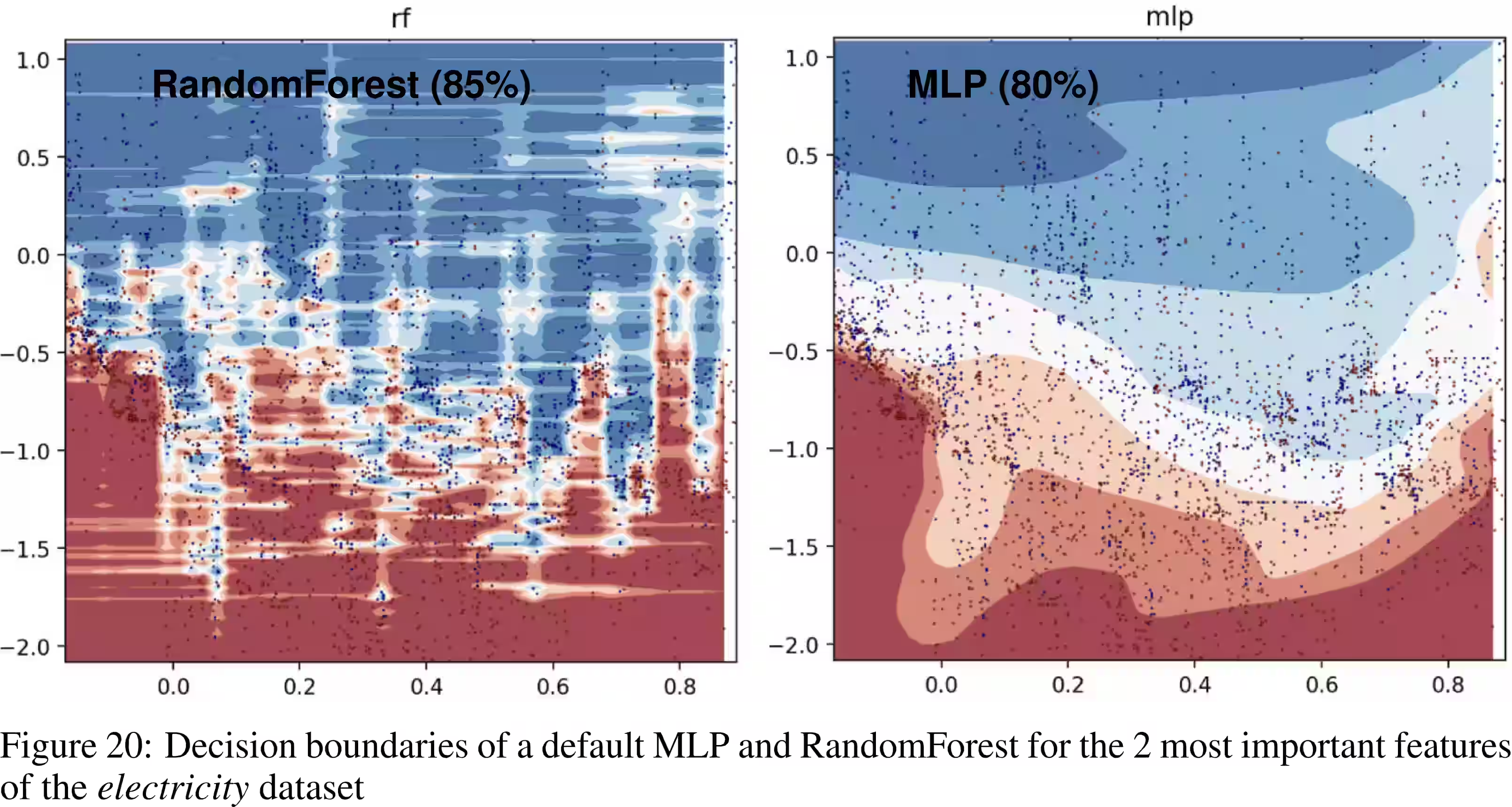
- They seem to be SOTA, or near-SOTA on some interesting challenging problems, notably tabular data (Gorishniy et al. 2023; Grinsztajn, Oyallon, and Varoquaux 2022).
Related: model averaging, neural ensembles, dropout, bootstrap.
1 Random trees, forests, jungles
- Awesome Random Forests
- how to do machine vision using random forests brought to you by the folks behind Kinect.
2 “Generalized” random forests
generalized random forests (Athey, Tibshirani, and Wager 2019) (implementation) describe themselves:
GRF extends the idea of a classic random forest to allow for estimating other statistical quantities besides the expected outcome. Each forest type, for example
quantile_forest, trains a random forest targeted at a particular problem, like quantile estimation. The most common use of GRF is in estimating treatment effects through the functioncausal_forest.
3 Self-regularizing properties
Jeremy Kun: Why Boosting Doesn’t Overfit:
Boosting, which we covered in gruesome detail previously, has a natural measure of complexity represented by the number of rounds you run the algorithm for. Each round adds one additional “weak learner” weighted vote. So running for a thousand rounds gives a vote of a thousand weak learners. Despite this, boosting doesn’t over-fit on many datasets. In fact, and this is a shocking fact, researchers observed that Boosting would hit zero training error, they kept running it for more rounds, and the generalization error kept going down! It seemed like the complexity could grow arbitrarily without penalty. […] this phenomenon is a fact about voting schemes, not boosting in particular.
🏗
4 Gradient boosting
The idea of gradient boosting originated in the observation by Leo Breiman that boosting can be interpreted as an optimization algorithm on a suitable cost function Breiman (1997). Explicit regression gradient boosting algorithms were subsequently developed by Jerome H. Friedman, (J. H. Friedman 2001, 2002) simultaneously with the more general functional gradient boosting perspective of Llew Mason, Jonathan Baxter, Peter Bartlett and Marcus Frean (Mason et al. 1999). The later two papers introduced the view of boosting algorithms as iterative functional gradient descent algorithms. That is, algorithms that optimize a cost function over function space by iteratively choosing a function (weak hypothesis) that points in the negative gradient direction. This functional gradient view of boosting has led to the development of boosting algorithms in many areas of machine learning and statistics beyond regression and classification.
5 Bayes
The Bayesian Additive Regression Trees Chipman, George, and McCulloch (2010), are wildly popular and successful in machine learning competitions. Kenneth Tay introduces them well.
6 Implementations.
6.1 LightGBM
LightGBM is a gradient boosting framework that uses tree-based learning algorithms. It is designed to be distributed and efficient with the following advantages:
- Faster training speed and higher efficiency.
- Lower memory usage.
- Better accuracy.
- Support of parallel, distributed, and GPU learning.
- Capable of handling large-scale data.
6.2 xgboost
XGBoost is an optimized distributed gradient boosting library designed to be highly efficient, flexible, and portable. It implements machine learning algorithms under the Gradient Boosting framework. XGBoost provides a parallel tree boosting (also known as GBDT, GBM) that solve many data science problems in a fast and accurate way. The same code runs on major distributed environments (Hadoop, SGE, MPI) and can solve problems beyond billions of examples.
See also chengsoonong/xgboost-tuner: A library for automatically tuning XGBoost parameters.
6.3 catboost
6.4 surfin
This R package computes uncertainty for random forest predictions using a fast implementation of random forests in C++. This is an exciting time for research into the theoretical properties of random forests. This R package aims to provide all state-of-the-art variance estimates in one place, to expedite research in this area and make it easier for practitioners to compare estimates.
Two variance estimates are provided: U-statistics based (Mentch & Hooker, 2016) and infinitesimal jackknife on bootstrap samples (Wager, Hastie, Efron, 2014), the latter as a wrapper to the authors’ R code randomForestCI.
More variance estimates coming soon: (1) Bootstrap-of-little-bags (Sexton and Laake 2009) (2) Infinitesimal jackknife on subsamples (Wager & Athey, 2017; Athey, Tibshirani, Wager, 2016) as a wrapper to the authors’ R package grf.
6.5 bartmachine
We present a new package in R implementing Bayesian additive regression trees (BART). The package introduces many new features for data analysis using BART such as variable selection, interaction detection, model diagnostic plots, incorporation of missing data and the ability to save trees for future prediction. It is significantly faster than the current R implementation, parallelized, and capable of handling both large sample sizes and high-dimensional data.