Garbled highlights from NeurIPS 2021
2021-11-05 — 2021-12-15
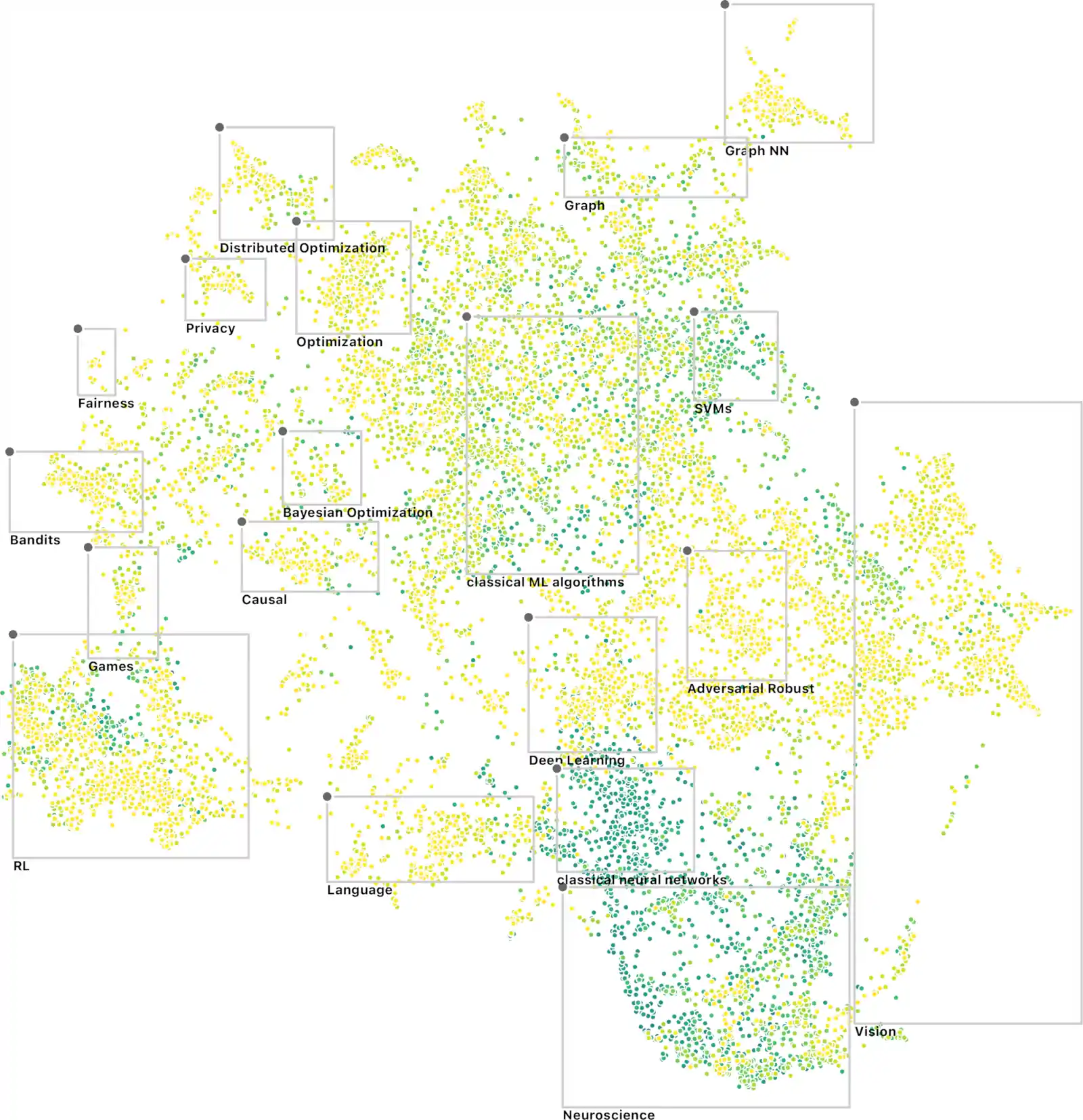
Wherein Workshops on Machine Learning for the Physical Sciences Are Catalogued, an Online Variational Filtering Method and a Laplace PyTorch Library Are Reported, and a NeurIPS Paper-Visualization Is Noted.
1 Workshops
1.1 Machine learning and the physical sciences
Three workshops on this theme with lots of overlap in content and speakers:
- Machine Learning and the Physical Sciences / Machine Learning and the Physical Sciences, NeurIPS 2021
- Physical Reasoning and Inductive Biases for the Real World / Physical Reasoning and Inductive Biases for the Real World
- The Symbiosis of Deep Learning and Differential Equations / The Symbiosis of Deep Learning and Differential Equations (DLDE)
1.2 Causal inference
1.3 Bayesian Deep learning
1.4 Strategic behaviour
1.5 DLDE
1.6 Physical Reasoning and Inductive Biases for the Real World
2 Tutorials
2.1 GPs
To be filed under GP regression.
2.2 Message passing
To be filed under message passing. There were some really nice observations by Wee Sun Lee about how to integrate neural function approximation into distributed inference.
2.3 Physics informed Deep Learning
To be filed under ml for physical sciences.
2.4 Real time optimisation for control
I am interested in this because it is dual to my interest in state estimation.
3 Interesting people and organizations encountered
Ekhi Ajuria Illarramendi, on solving the Poisson equation
4 Interesting papers
These were papers I encountered during Neurips
not all were published in Neurips.
Campbell et al. (2021)
We present a variational method for online state estimation and parameter learning in state-space models (SSMs), a ubiquitous class of latent variable models for sequential data. As per standard batch variational techniques, we use stochastic gradients to simultaneously optimize a lower bound on the log evidence with respect to both model parameters and a variational approximation of the states’posterior distribution. However, unlike existing approaches, our method is able to operate in an entirely online manner, such that historic observations do not require revisitation after being incorporated and the cost of updates at each time step remains constant, despite the growing dimensionality of the joint posterior distribution of the states. This is achieved by utilising backward decompositions of this joint posterior distribution and of its variational approximation, combined with Bellman-type recursions for the evidence lower bound and its gradients. We demonstrate the performance of this methodology across several examples, including high-dimensional SSMs and sequential Variational Auto-Encoders.
This is an amazing trick in system identification and variational state filters at once.
- andrew-cr/online_var_fil: Code for our paper: Online Variational Filtering and Parameter Learning
- Online Variational Filtering and Parameter Learning
Daxberger et al. (2021)
Bayesian formulations of deep learning have been shown to have compelling theoretical properties and offer practical functional benefits, such as improved predictive uncertainty quantification and model selection. The Laplace approximation (LA) is a classic, and arguably the simplest family of approximations for the intractable posteriors of deep neural networks. Yet, despite its simplicity, the LA is not as popular as alternatives like variational Bayes or deep ensembles. This may be due to assumptions that the LA is expensive due to the involved Hessian computation, that it is difficult to implement, or that it yields inferior results. In this work we show that these are misconceptions: we (i) review the range of variants of the LA including versions with minimal cost overhead; (ii) introduce
laplace, an easy-to-use software library for PyTorch offering user-friendly access to all major flavors of the LA; and (iii) demonstrate through extensive experiments that the LA is competitive with more popular alternatives in terms of performance, while excelling in terms of computational cost. We hope that this work will serve as a catalyst to a wider adoption of the LA in practical deep learning, including in domains where Bayesian approaches are not typically considered at the moment.
https://github.com/AlexImmer/Laplace
Galimberti et al. (2021)
Huang, Bai, and Kolter (2021)
Abstract: Recent research in deep learning has investigated two very different forms of ‘’implicitness’’: implicit representations model high-frequency data such as images or 3D shapes directly via a low-dimensional neural network (often using e.g., sinusoidal bases or nonlinearities); implicit layers, in contrast, refer to techniques where the forward pass of a network is computed via non-linear dynamical systems, such as fixed-point or differential equation solutions, with the backward pass computed via the implicit function theorem. In this work, we demonstrate that these two seemingly orthogonal concepts are remarkably well-suited for each other. In particular, we show that by exploiting fixed-point implicit layer to model implicit representations, we can substantially improve upon the performance of the conventional explicit-layer-based approach. Additionally, as implicit representation networks are typically trained in large-batch settings, we propose to leverage the property of implicit layers to amortize the cost of fixed-point forward/backward passes over training steps — thereby addressing one of the primary challenges with implicit layers (that many iterations are required for the black-box fixed-point solvers). We empirically evaluated our method on learning multiple implicit representations for images, videos and audios, showing that our \((\textrm{Implicit})^2\) approach substantially improve upon existing models while being both faster to train and much more memory efficient.
Kawar, Vaksman, and Elad (2021)
Lee and Bareinboim (2021)
Causal effect identification is concerned with determining whether a causal effect is computable from a combination of qualitative assumptions about the underlying system (e.g., a causal graph) and distributions collected from this system. Many identification algorithms exclusively rely on graphical criteria made of a non-trivial combination of probability axioms, do-calculus, and refined c-factorization (e.g., Lee & Bareinboim, 2020). In a sequence of increasingly sophisticated results, it has been shown how proxy variables can be used to identify certain effects that would not be otherwise recoverable in challenging scenarios through solving matrix equations (e.g., Kuroki & Pearl, 2014; Miao et al., 2018). In this paper, we develop a new causal identification algorithm which utilizes both graphical criteria and matrix equations. Specifically, we first characterize the relationships between certain graphically-driven formulae and matrix multiplications. With such characterizations, we broaden the spectrum of proxy variable based identification conditions and further propose novel intermediary criteria based on the pseudoinverse of a matrix. Finally, we devise a causal effect identification algorithm, which accepts as input a collection of marginal, conditional, and interventional distributions, integrating enriched matrix-based criteria into a graphical identification approach.
Lorch et al. (2021)
Bayesian structure learning allows inferring Bayesian network structure from data while reasoning about the epistemic uncertainty — a key element towards enabling active causal discovery and designing interventions in real world systems. In this work, we propose a general, fully differentiable framework for Bayesian structure learning (DiBS) that operates in the continuous space of a latent probabilistic graph representation. Contrary to existing work, DiBS is agnostic to the form of the local conditional distributions and allows for joint posterior inference of both the graph structure and the conditional distribution parameters. This makes our formulation directly applicable to posterior inference of nonstandard Bayesian network models, e.g., with nonlinear dependencies encoded by neural networks. Using DiBS, we devise an efficient, general purpose variational inference method for approximating distributions over structural models. In evaluations on simulated and real-world data, our method significantly outperforms related approaches to joint posterior inference.
Lütjens et al. (2021)
Meronen, Trapp, and Solin (2021)
Neural network models are known to reinforce hidden data biases, making them unreliable and difficult to interpret. We seek to build models that `know what they do not know’ by introducing inductive biases in the function space. We show that periodic activation functions in Bayesian neural networks establish a connection between the prior on the network weights and translation-invariant, stationary Gaussian process priors. Furthermore, we show that this link goes beyond sinusoidal (Fourier) activations by also covering triangular wave and periodic ReLU activation functions. In a series of experiments, we show that periodic activation functions obtain comparable performance for in-domain data and capture sensitivity to perturbed inputs in deep neural networks for out-of-domain detection.
Nonnenmacher and Greenberg (2021)
Norcliffe et al. (2021)
van Krieken, Tomczak, and Teije (2021)
Stochastic AD extends AD to stochastic computation graphs with sampling steps, which arise when modelers handle the intractable expectations common in Reinforcement Learning and Variational Inference. However, current methods for stochastic AD are limited: They are either only applicable to continuous random variables and differentiable functions, or can only use simple but high variance score-function estimators. To overcome these limitations, we introduce Storchastic, a new framework for AD of stochastic computation graphs. Storchastic allows the modeler to choose from a wide variety of gradient estimation methods at each sampling step, to optimally reduce the variance of the gradient estimates. Furthermore, Storchastic is provably unbiased for estimation of any-order gradients, and generalizes variance reduction techniques to higher-order gradient estimates. Finally, we implement Storchastic as a PyTorch library at HEmile/storchastic.
Adam et al. (2021)
Bubeck and Sellke (2021)